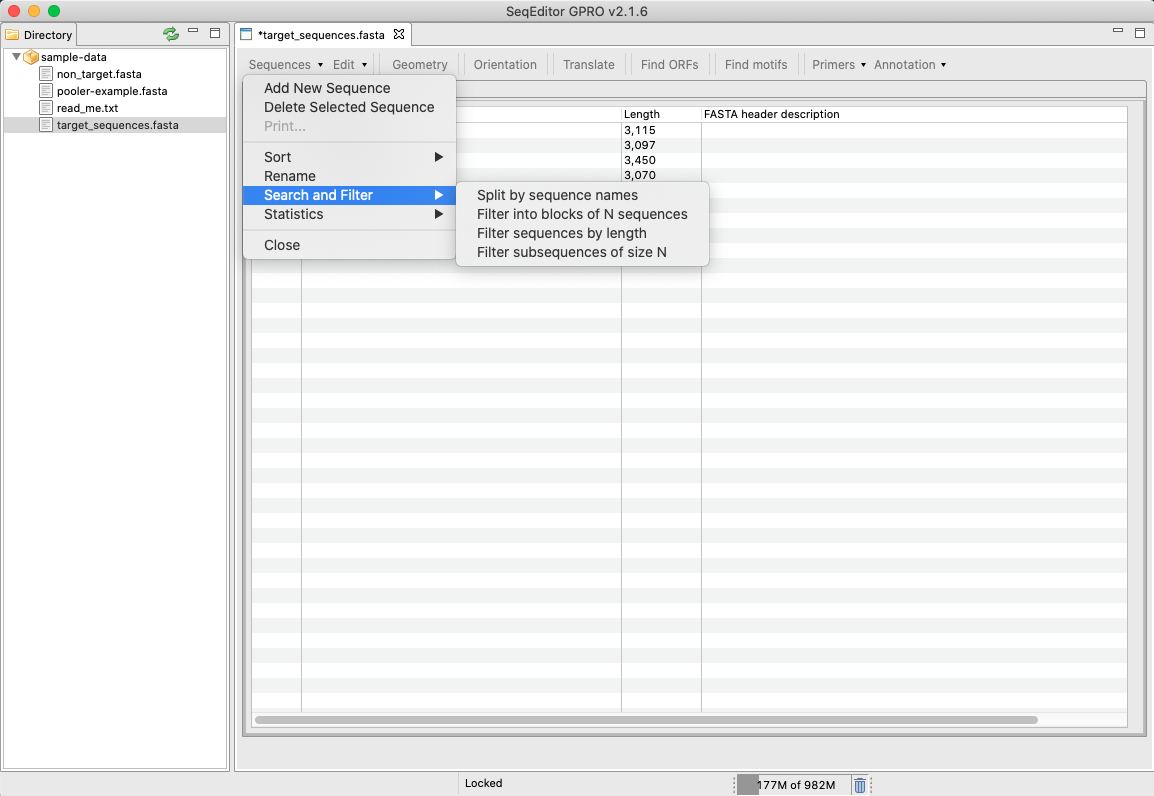
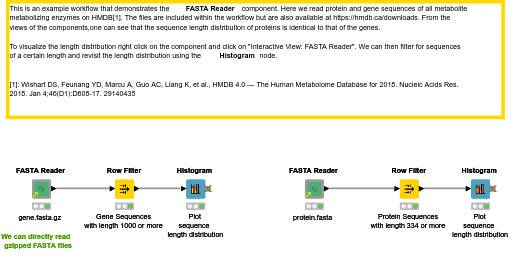
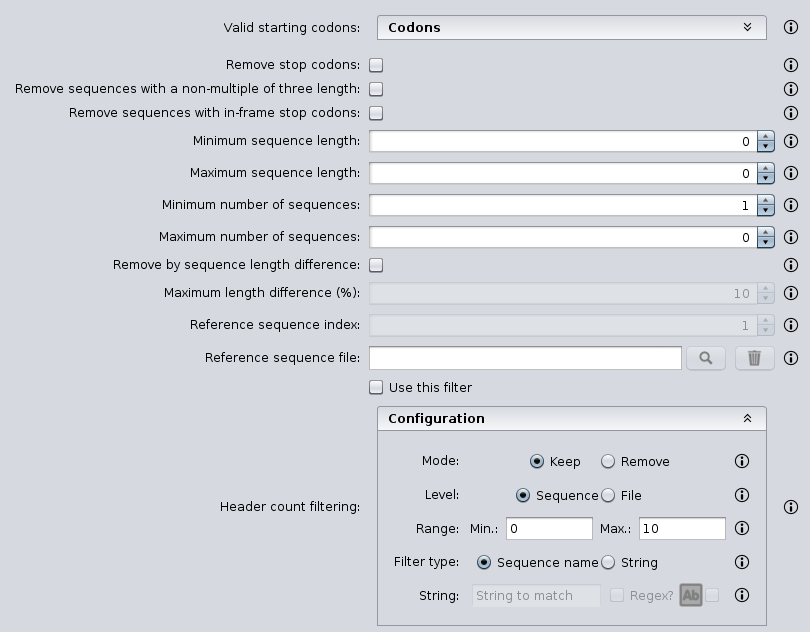
Annotation for de novo Assembled Seq. Contents 1 FASTA File Length Filter 2 Gene Prediction Choose GENSCAN or GeneMark.hmm 3 Gene Prediction to FASTA Converts GENSCAN or GeneMark.hmm output file to FASTA format 4 transcriptsToOrfs Trinity ...
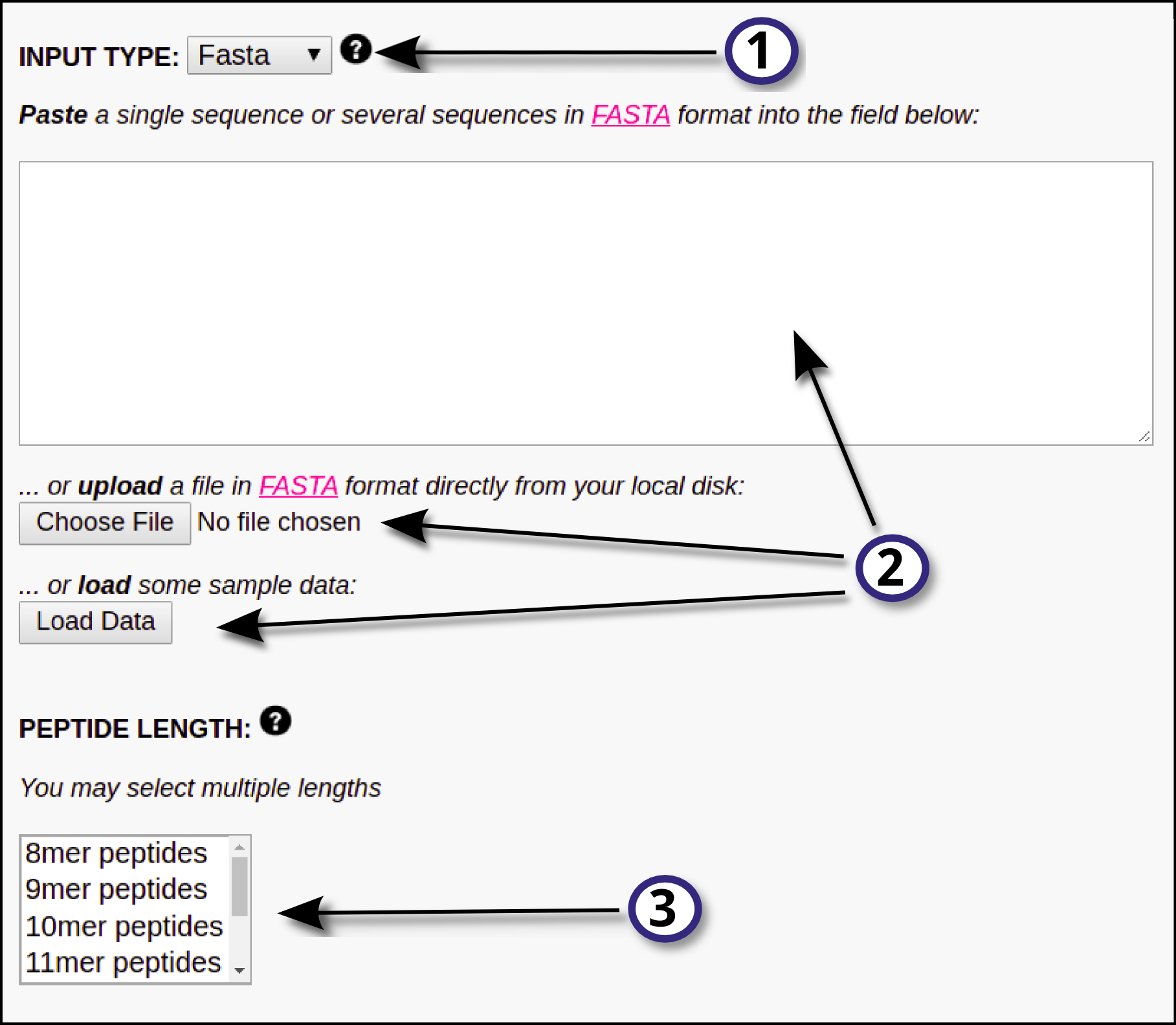
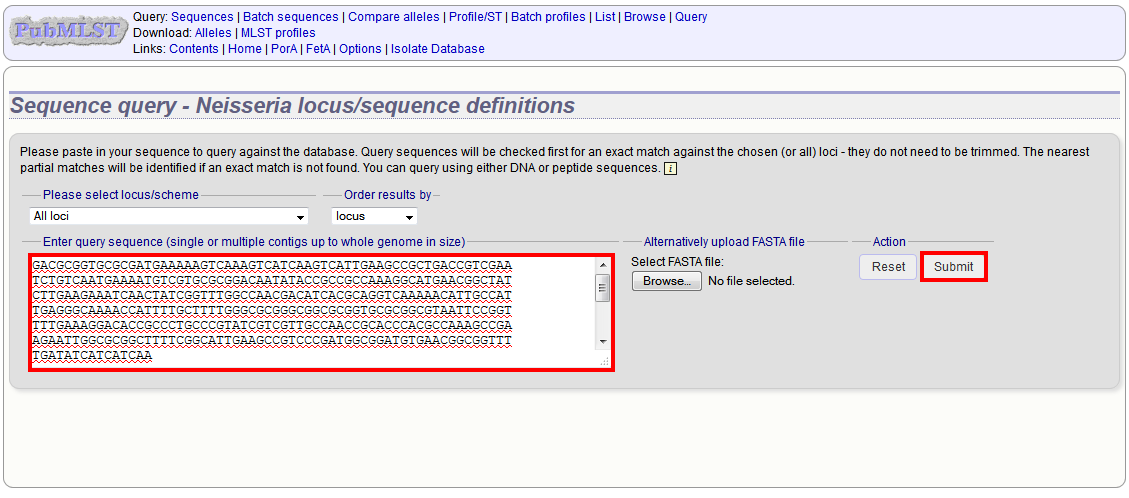
INPUT DATA In this section, the user must define the input for the prediction server following these steps: 1) Specify the desired type of input data (FASTA or PEPTIDE) using the drop down menu. 2) Provide the input data by means of pasting the data into the ...